
Till date communicable disease control relies on effective structures, including highly developed surveillance systems. An integrated approach to the surveillance of communicable disease and infectious agents requires harmonization of molecular typing activities across all sectors (one health approach). Sharing molecular- and epidemiologic-data prospectively is as important as it is significant.
Next generation sequencing (NGS)-based molecular typing methods are in the process of revolutionizing public health, food, and veterinary microbiology. Whole genome sequencing (WGS) data produced by NGS are information-rich and provide an essentially unlimited level of data, such as resistance and virulence markers or strain relationship, that is ‘hypothesis-free’ in nature, thereby enabling new and so far unrecognized observations to be made. WGS is a break-through ‘generalist’ approach that ‘fits all’ pathogens (bacteria, viruses, fungi, or parasites) and study types (i.e., diagnostics, early warning and response system [EWRS], outbreak investigation, risk potential assessment, source attribution, and population genetics).
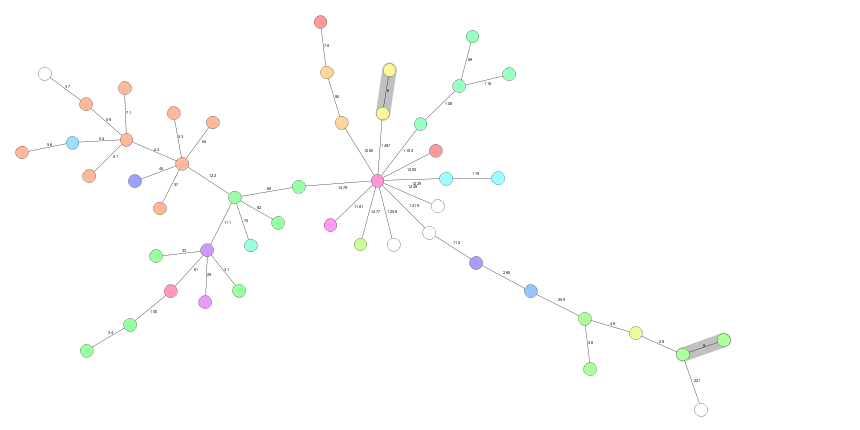
Therefore, the Microbial Genomic Surveillance (miGenomeSurv) consortium aims to support a nation-wide, prospective integrated genomic surveillance for recognized high-priority organisms, such as enterohemorrhagic Escherichia coli (EHEC), Listeria monocytogenes, multidrug-resistant (MDR) Mycobacterium tuberculosis, and vancomycin-resistant Enterococci (VRE). In this manner a cross-sectorial early detection of clusters is possible, and consumer and patient safety is improved by avoiding or at least containing the spread of infectious disease.
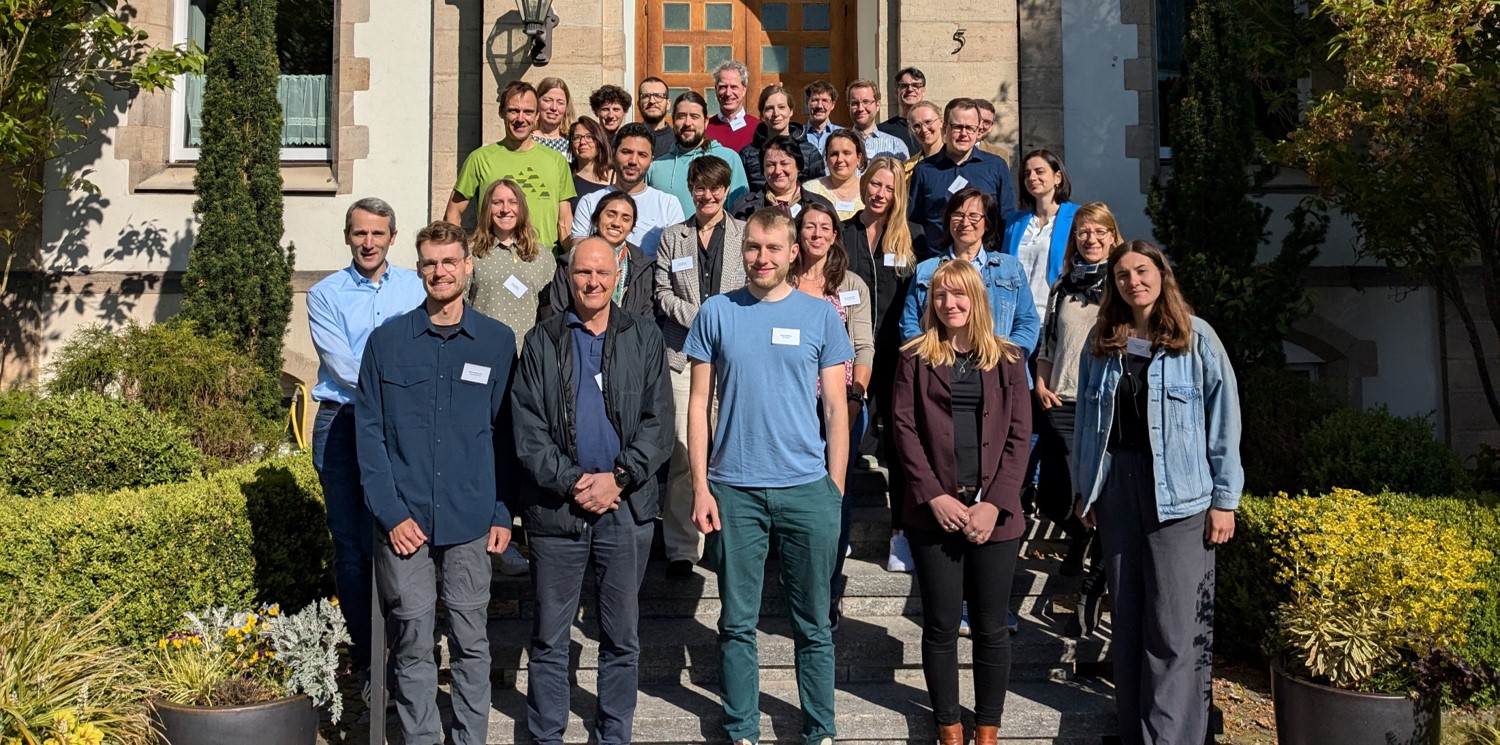
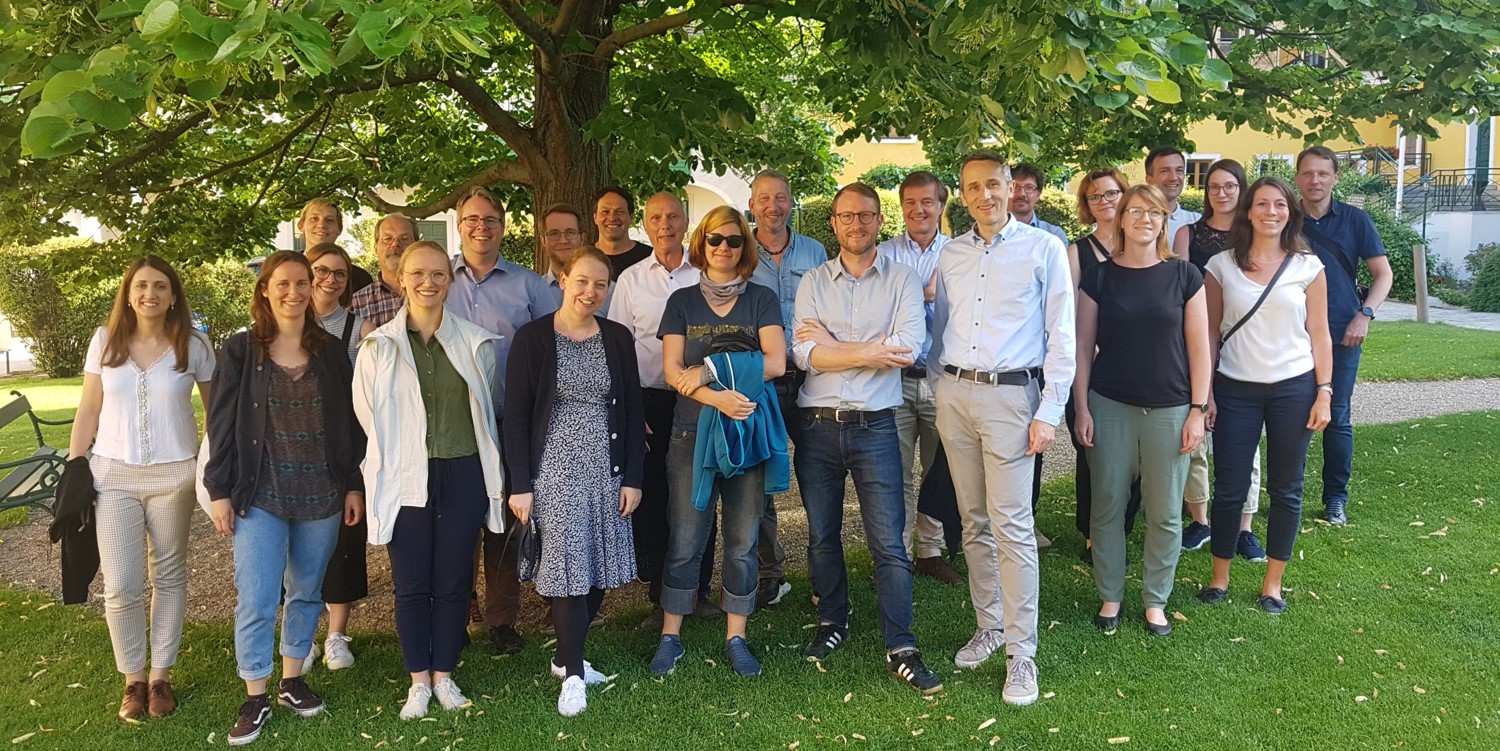
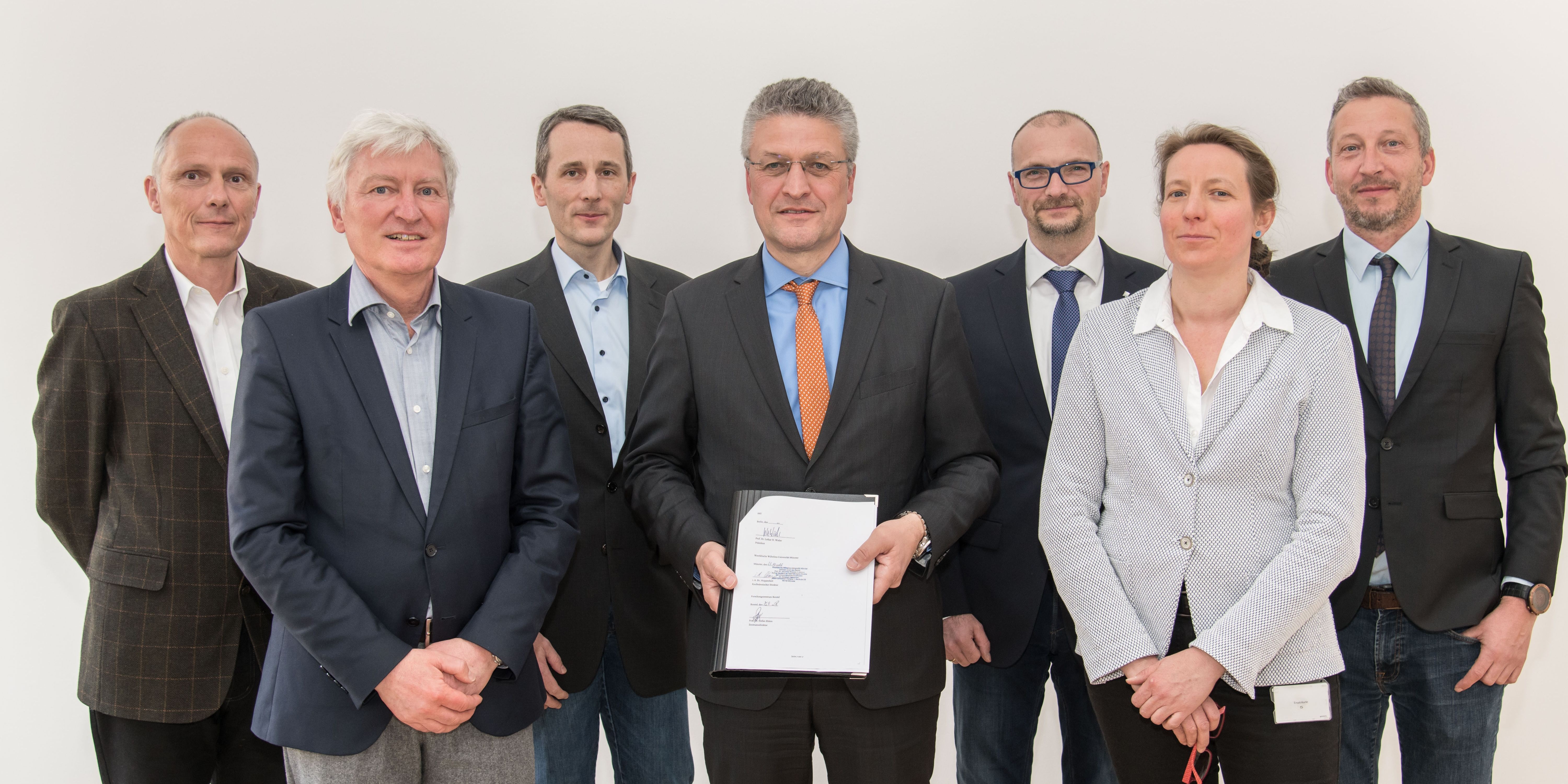
Current network